This is a default text for your page Johnson's Monday Lab Sandbox for Insulin Receptor. Click above on edit this page to modify. Be careful with the < and > signs.
You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
Function of the Receptor
The insulin receptor resides within the plasma membrane of insulin target cells of different organs, such as the liver, and tissues including skeletal muscle and adipose. Activation of the insulin receptor is dependent upon insulin binding. Once activated, the receptor serves as the gateway for the regulation of various cellular processes. These processes include but are not limited to glucose transport, glycogen storage, autophagy, apoptosis, and gene expression. Additionally, the insulin receptor has been associated with the development of diseases such as Alzheimer's, Type II Diabetes, and cancer [3]. Characterization of the structure of the insulin receptor as well as understanding of the molecular mechanisms which initiate a conformational change are important for understanding the role that the insulin receptor plays within a cell and in the development of disease.
Insulin
WHAT IS INSULIN. WHAT DOES IT LOOK LIKE? WHERE IS IT MADE? WHAT DOES IT DO? WHERE DOES IT GO? WHY DO WE NEED IT?
Insulin is a horomone that is synthesized and secreted from the pancreas in response to high concentrations of glucose in the blood. Once it is secreted, it will move through the blood stream and attach to an insulin receptor. Once multiple insulins are bound to the receptor, it is activated and as mentioned previously, the regulation of various cellular processes is initiated.
Structure
The insulin receptor is a receptor tyrosine kinase. It is a heterotetramer which is constructed from two homodimers. Each homodimer maintains an extracellular domain, transmembrane helix, and an intracellular domain. The extracellular domain is divided into alpha and beta subunits. The alpha subunit is characterized by two leucine-rich regions and one cysteine rich region. The beta subunit contains three fibronectin type III domains. The alpha and beta subunits of the extracellular domains fold over one another and form a "V" shape when the insulin receptor is unactivated. Upon activation, the extracellular domain undergoes a conformational change and forms a "T" shape.
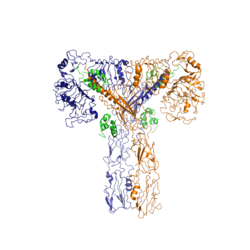
Figure 1: Insulin receptor in the "T" shape active conformation with four insulins bound
An additional component to the ectodomain is the alpha chain C-terminal helix [4]. The alpha-CT is a single alpha helix and it plays an important role in insulin binding and stabilization of the "T" shape activated conformation. The alpha-CT interacts with a leucine rich region of the alpha subunit and a fibronectin type III region of the beta subunit to form an insulin binding site [4].
The structure of the extracellular domain is stabilized through covalent bonds. The alpha subunits are linked through two disulfide bonds. Cys468 and Cys524 of one alpha subunit are bound to Cys435 and Cys524 of the other alpha subunit, respectively [5].
The insulin receptor extends intracellularly from the beta subunits of the ectodomain by way of a transmembrane helix. Intracellularly, the insulin receptor contains two tyrosine kinase domains.
Insulin Binding
The insulin receptor unit has four separate sites for the insulin molecule to bind to. There are two pairs of two identical binding sites referred to as 1 and 1' and then 2 and 2'. The insulin molecules bind to these sites mostly through hydrophobic interactions. Despite a majority of the interactions being similar, sites 1 and 1' have a higher binding affinity than sites 2 and 2' due to site one having a larger surface area (706 square angstroms) exposed for insulin to bind to compared to site 2 (394 square angstroms)[4].
It was found that at least three insulin molecules would have to bind to the receptor for the receptor to take on its active “T-state” conformation [4]. The difference between the fully bound state with four insulins and the three insulin bound state is minimal compared to the difference between two and three insulins bound [4].
The insulin molecules in site 1 and 1' have their main interactions with an in the insulin receptor. The insulin molecules are shown in green and the insulin receptor is shown in orange. The insulin molecules in site 2 and 2' have their main interactions with the residues that comprise some of the of the insulin receptor. The red molecules are insulin and the yellow is the beta sheets of the insulin receptor.
Conformational Changes
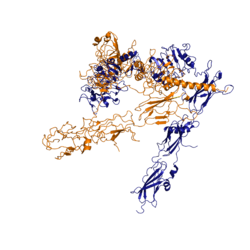
Figure 2: Conformational change of insulin receptor protomer from inactive (blue) to active (orange) form upon insulin binding.
The conformational change between the inverted "V" shape and the "T" shape of the insulin receptor is induced by insulin binding. When an insulin molecule binds to site 1 of the alpha subunit, the respective protomer is recruited and a slight inward movement of the fibronectin type III domains of the beta subunit is initiated. Binding of insulin to both protomers establishes a full activation of the insulin receptor. This activation is demonstrated through the inward movement of both protomers. This motion has been referred to as a "hinge" motion [4] as both protomers "swing" in towards one another.
As the fibronectin type III domains of the beta subunit swing inward, the alpha subunits also undergo a conformational change upon insulin binding. As insulin binds to site 1, the leucine rich region of one protomer interacts with the alpha-CT and the FNIII-1 domains of the other protomer to form a binding site. These interactions are referred to as a tripartite interface [4]. In order for the tripartite interface to form, the alpha subunits of each protomer must undergo a "folding" motion.
While there is an explanation for which conformational changes of the insulin receptor take place, there is no explanation for mechanism by which the conformational changes are executed [4].
Type II Diabetes
WHAT IS T2D? HOW DOES IT RELATE TO THE INSULIN RECEPTOR? WHAT DOES A NON T2D SYSTEM LOOK LIKE? HOW DOES IT COMPARE TO T1D? WHY IS THIS IMPORTANT?
References
[6]
[7]
[3]
[5]
[4]
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ 3.0 3.1 Scapin G, Dandey VP, Zhang Z, Prosise W, Hruza A, Kelly T, Mayhood T, Strickland C, Potter CS, Carragher B. Structure of the Insulin Receptor-Insulin Complex by Single Particle CryoEM analysis. Nature. 2018 Feb 28. pii: nature26153. doi: 10.1038/nature26153. PMID:29512653 doi:http://dx.doi.org/10.1038/nature26153
- ↑ 4.0 4.1 4.2 4.3 4.4 4.5 4.6 4.7 4.8 Uchikawa E, Choi E, Shang G, Yu H, Bai XC. Activation mechanism of the insulin receptor revealed by cryo-EM structure of the fully liganded receptor-ligand complex. Elife. 2019 Aug 22;8. pii: 48630. doi: 10.7554/eLife.48630. PMID:31436533 doi:http://dx.doi.org/10.7554/eLife.48630
- ↑ 5.0 5.1 Schaffer L, Ljungqvist L. Identification of a disulfide bridge connecting the alpha-subunits of the extracellular domain of the insulin receptor. Biochem Biophys Res Commun. 1992 Dec 15;189(2):650-3. PMID:1472036
- ↑ Boucher J, Kleinridders A, Kahn CR. Insulin receptor signaling in normal and insulin-resistant states. Cold Spring Harb Perspect Biol. 2014 Jan 1;6(1). pii: 6/1/a009191. doi:, 10.1101/cshperspect.a009191. PMID:24384568 doi:http://dx.doi.org/10.1101/cshperspect.a009191
- ↑ De Meyts P. The structural basis of insulin and insulin-like growth factor-I receptor binding and negative co-operativity, and its relevance to mitogenic versus metabolic signalling. Diabetologia. 1994 Sep;37 Suppl 2:S135-48. doi: 10.1007/bf00400837. PMID:7821729 doi:http://dx.doi.org/10.1007/bf00400837
Student Contributors
- Maxwell Todd
- Abby Hillan
- Andrew Scheel


