This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Daniel Mulawa/Sandbox 1
From Proteopedia
| Line 25: | Line 25: | ||
===Substrate Structure=== | ===Substrate Structure=== | ||
Though gamma secretase has multiple substrates, the substrate of main concern is called Amyloid Precursor Protein (APP). APP is composed of an N-terminal loop, a transmembrane (TM) helix, and a C-terminal β-strand. The uses lateral diffusion as a mechanism of entry into the enzyme, and once in place, the TM helix is anchored by hydrogen bonds. In order to differentiate substrates, the β-strand is often the main point of identification for the enzyme. After this, the helix undergoes unwinding and the process of catalysis can begin. | Though gamma secretase has multiple substrates, the substrate of main concern is called Amyloid Precursor Protein (APP). APP is composed of an N-terminal loop, a transmembrane (TM) helix, and a C-terminal β-strand. The uses lateral diffusion as a mechanism of entry into the enzyme, and once in place, the TM helix is anchored by hydrogen bonds. In order to differentiate substrates, the β-strand is often the main point of identification for the enzyme. After this, the helix undergoes unwinding and the process of catalysis can begin. | ||
| - | TM helix anchored by hydrogen bonds | ||
| - | Unwinding model | ||
| - | used for substrate recognition | ||
===Lid Complex=== | ===Lid Complex=== | ||
Revision as of 20:37, 24 March 2020
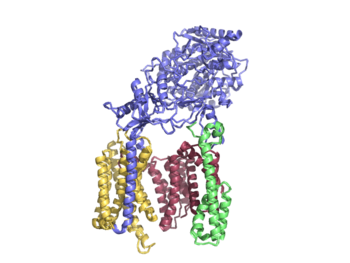
Gamma Secretase
=Human Gamma Secretase=
| |||||||||||
References
1.Bai X, Yan C, Guanghui Yang, et al. 2015. An atomic structure of human γ-secretase. Nature. 525:212-217. 2.Carroll CM, and Li YM. 2016. Physiological and pathological roles of the γ-secretase complex. Brain research bulletin. 126:199-206. 3.Yang G, Zhou R, Shi Y. 2017. Cryo-EM structures of human γ-secretase. Current Opinion in Structural Biology. 46:55–64. 4.Yang G, Zhou R, Zhou Q, et al. 2019. Structural basis of Notch recognition by human γ-secretase. Nature. 565: 192-197. 5.Zhou R, Yang G, Guo X, et al. 2019. Recognition of the amyloid precursor protein by human γ-secretase. Science. 363:1-8.
Student Contributors
Layla Wisser Daniel Mulawa