This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:George G. Papadeas/Sandbox VKOR
From Proteopedia
| Line 6: | Line 6: | ||
== Introduction== | == Introduction== | ||
=== Biological Role === | === Biological Role === | ||
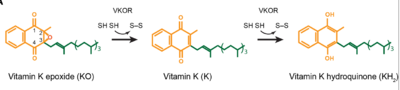
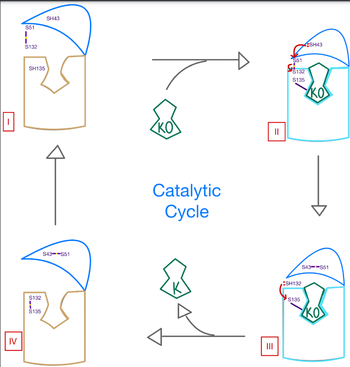
| - | <scene name='90/906893/Vkor_structure/1'>Vitamin K epoxide reductase</scene> (VKOR) is a reducing enzyme composed of 4-helices that spans the endoplasmic reticulum as a transmembrane protein <ref>DOI 10.1126</ref>. Its enzymatic role is reducing <scene name='90/906893/Vkor_with_ko/1'>vitamin K epoxide</scene> (KO) to Vitamin K Hydroquinone (KH2) <ref>DOI 10.1021</ref> (Figure 1). The mechanism first occurs through the binding KO and using two cysteine residues to reduce KO into [https://en.wikipedia.org/wiki/Vitamin_K Vitamin K]. Then, a second pair of cysteine residues will reduce Vitamin K into the final product, KH2 (Figure 1). One of VKORs primary roles is to assist in blood coagulation through this KH2 regeneration mechanism.[[Image:VKOR_mechanism_2D.png|400 px|right|thumb|Figure 1. Mechanism of KO oxidation into KH2.]] With Vitamin K as a cofactor, the [https://www.britannica.com/science/bleeding/The-extrinsic-pathway-of-blood-coagulation#ref64617 γ-carboxylase] enzyme will enact post-translational modification on KH2, oxidizing it back to KO. The oxidation of KH2 by γ-carboxylase is coupled with the carboxylation of a glutamate residue to form γ-carboxyglutamate. The coupling of this oxidation and carboxylation is necessary for the activation of several clotting factors in the coagulation cascade. | + | <scene name='90/906893/Vkor_structure/1'>Vitamin K epoxide reductase</scene> (VKOR) is a reducing enzyme composed of 4-helices that spans the endoplasmic reticulum as a transmembrane protein <ref>DOI 10.1126</ref>. Its enzymatic role is reducing <scene name='90/906893/Vkor_with_ko/1'>vitamin K epoxide</scene> (KO) to Vitamin K Hydroquinone (KH2) <ref>DOI 10.1021</ref> (Figure 1). The mechanism first occurs through the binding KO and using two cysteine residues to reduce KO into [https://en.wikipedia.org/wiki/Vitamin_K Vitamin K]. Then, a second pair of cysteine residues will reduce Vitamin K into the final product, KH2 (Figure 1). One of VKORs primary roles is to assist in blood coagulation through this KH2 regeneration mechanism.[[Image:VKOR_mechanism_2D.png|400 px|right|thumb|Figure 1. Mechanism of KO oxidation into KH2<ref>DOI 10.1126</ref>.]] With Vitamin K as a cofactor, the [https://www.britannica.com/science/bleeding/The-extrinsic-pathway-of-blood-coagulation#ref64617 γ-carboxylase] enzyme will enact post-translational modification on KH2, oxidizing it back to KO. The oxidation of KH2 by γ-carboxylase is coupled with the carboxylation of a glutamate residue to form γ-carboxyglutamate. The coupling of this oxidation and carboxylation is necessary for the activation of several clotting factors in the coagulation cascade. |
=== Author's Notes === | === Author's Notes === | ||
Revision as of 23:54, 18 April 2022
VKOR
| |||||||||||
References
1. DJin, Da-Yun, Tie, Jian-Ke, and Stafford, Darrel W. "The Conversion of Vitamin K Epoxide to Vitamin K Quinone and Vitamin K Quinone to Vitamin K Hydroquinone Uses the Same Active Site Cysteines." Biochemistry 2007 46 (24), 7279-7283 [1].
2. Li, Weikai et al. “Structure of a bacterial homologue of vitamin K epoxide reductase.” Nature vol. 463,7280 (2010): 507-12. doi:10.1038/nature08720.
3. Liu S, Li S, Shen G, Sukumar N, Krezel AM, Li W. Structural basis of antagonizing the vitamin K catalytic cycle for anticoagulation. Science. 2021 Jan 1;371(6524):eabc5667. doi: 10.1126/science.abc5667. Epub 2020 Nov 5. PMID: 33154105; PMCID: PMC7946407.
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
- ↑ Unknown PubmedID 10.1126
- ↑ Unknown PubmedID 10.1021
- ↑ Unknown PubmedID 10.1126
- ↑ Unknown PubmedID 10.1126