This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
BASIL2022GV3R8E
From Proteopedia
| Line 14: | Line 14: | ||
[[Image:DALI.png]] | [[Image:DALI.png]] | ||
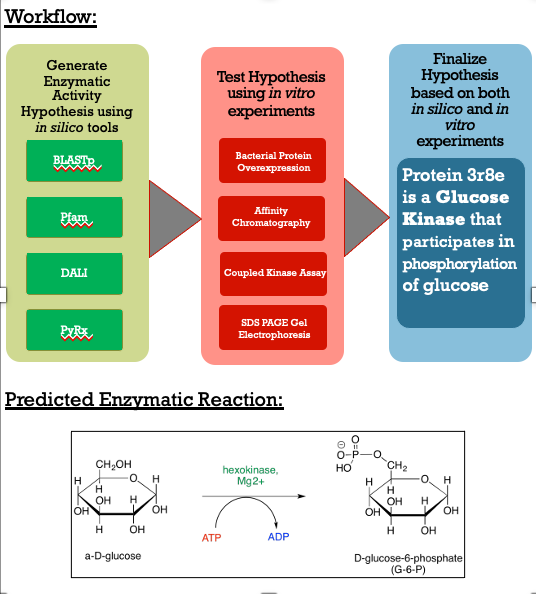
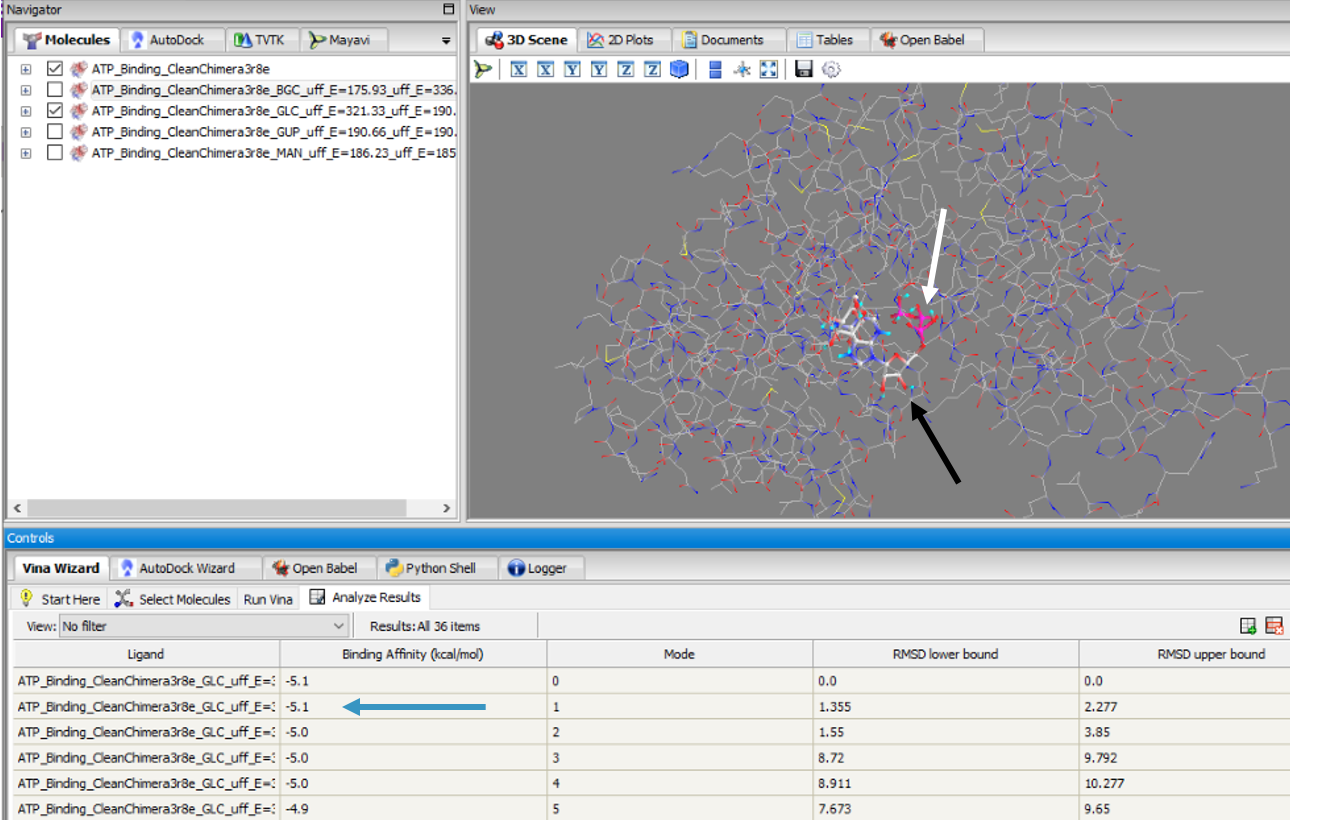
These results along with supporting evidence from the other in silico tools allowed us to then conduct in silico docking experiments to further confirm our hypothesis. Below is a figure with the PyRx docking results of glucose and ATP with our protein. | These results along with supporting evidence from the other in silico tools allowed us to then conduct in silico docking experiments to further confirm our hypothesis. Below is a figure with the PyRx docking results of glucose and ATP with our protein. | ||
| - | + | [[Image:PyRx.png]] | |
The -5.1 kcal/mol value shows that our protein of interest hypothesis of a glucose kinase is strong. The confidence behind our in silico results allowed us to move into testing our hypothesis in vitro. | The -5.1 kcal/mol value shows that our protein of interest hypothesis of a glucose kinase is strong. The confidence behind our in silico results allowed us to move into testing our hypothesis in vitro. | ||
== Experimental Results/Function == | == Experimental Results/Function == | ||
Revision as of 00:26, 21 April 2022
Your Heading Here (maybe something like 'Structure')
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644
3. Blastp [Internet]. Bethesda (MD): Natiobal Library of Medicine (US), National Center for Biotechnology Information; 2004- [cited 2022 March]. Available from: (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins)
4. BASIL. https://basilbiochem.github.io/basil/
5. Holm L (2020) Using Dali for protein structure comparison. Methods Mol. Biol. 2112, 29-42.
6. Small- Molecule Library Screening by Docking with PyRx. .Dallakyan S, Olson AJ Methods Mol Biol. 2015;1263:243-50. The full-text is available at https://www.researchgate.net/publications/2739554875. Small-Molecule Library Screening by Docking with PyRx.
7. Pfam: The Protein families database in 2021 J. Mistry, S. Chuguransky, L. Williams, M. Qureshi, G.A. Salazar, E.L.L. Sonnhammer, S.C.E. Tosatto, L. Paladin, S. Raj, L.J. Richardson, R.D. Finn, A. Bateman Nucleic Acids Research (2020) doi: 10.1093/nar/gkaa913
8. The PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC.
Proteopedia Page Contributors and Editors (what is this?)
Dalton Dencklau, Michel Evertsen, Bonnie Hall, Jaime Prilusky