We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
LiLac - a biosensor for Lactate
From Proteopedia
(Difference between revisions)
| Line 17: | Line 17: | ||
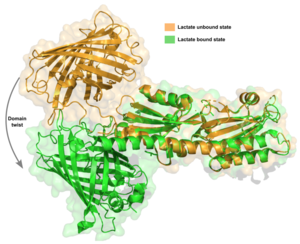
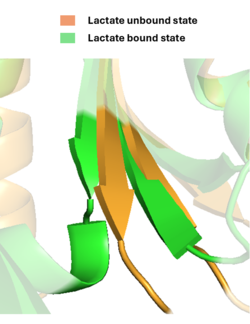
Second, a <scene name='10/1096830/B_l/1'>β-strand</scene> at the back of the binding pocket, near its very C-terminus, shortens by one amino acid to accommodate lactate when it binds. This shortening is associated with a retraction of the TlpC “tail” in LiLac, which protrudes in the lactate-free form. Third, the tail undergoes a remarkable transformation in response to lactate binding, curling up into a short helical turn. | Second, a <scene name='10/1096830/B_l/1'>β-strand</scene> at the back of the binding pocket, near its very C-terminus, shortens by one amino acid to accommodate lactate when it binds. This shortening is associated with a retraction of the TlpC “tail” in LiLac, which protrudes in the lactate-free form. Third, the tail undergoes a remarkable transformation in response to lactate binding, curling up into a short helical turn. | ||
| - | [[Image:Bturn.png| | + | [[Image:Bturn.png|250px|Conformational changes in the beta sheet domain]] |
| - | + | ||
---- | ---- | ||
| + | == '''Mechanism for decreased lifetime upon lactate binding''' == | ||
| - | + | The chromophore in our lactate-bound, low- lifetime structure lacked the seal normally seen in mTurquoise, almost certainly stabilizing the chromophore much less. In contrast, the chromophore in our lactate-free, high- lifetime structure was sealed shut. The “seal” for the mTurquoise portion of LiLac in a high-lifetime state was provided by the engineered <scene name='10/1096830/C_link/1'>C terminal linker</scene>, rather than the sequence that normally comprises the N-terminal half of β7; the N-terminal linker was largely disordered. The protein backbone of the C-terminal linker, as opposed to any of its specific amino acid side chains, is probably the “business end”. | |
| - | + | ||
| - | + | ||
| - | + | ||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Revision as of 20:28, 29 November 2025
INTRODUCTION TO A LACTATE SENSOR
| |||||||||||
References
- ↑ doi: https://dx.doi.org/10.1038/s41467-022-30685-x
- ↑ Machuca MA, Johnson KS, Liu YC, Steer DL, Ottemann KM, Roujeinikova A. Helicobacter pylori chemoreceptor TlpC mediates chemotaxis to lactate. Sci Rep. 2017 Oct 26;7(1):14089. doi: 10.1038/s41598-017-14372-2. PMID:29075010 doi:http://dx.doi.org/10.1038/s41598-017-14372-2
- ↑ Rosen PC, Horwitz SM, Brooks DJ, Kim E, Ambarian JA, Waidmann L, Davis KM, Yellen G. State-dependent motion of a genetically encoded fluorescent biosensor. Proc Natl Acad Sci U S A. 2025 Mar 11;122(10):e2426324122. PMID:40048274 doi:10.1073/pnas.2426324122