This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Resolution
From Proteopedia
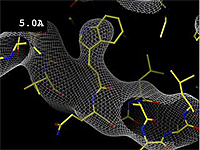
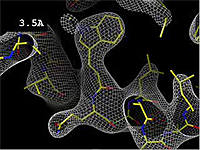
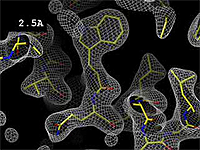
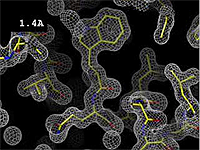
Resolution is an average value for the uncertainty of atomic positions in a crystallographic model. High values for resolution (e.g. 5.0 Å) mean high uncertainty, and low values (e.g. 1.0 Å) mean much less uncertainty. 2.05 Å is the median resolution for X-ray crystallographic results in the Protein Data Bank (43,066 on May 2, 2008).
The uncertainty for each atom is quantitated in its temperature value.
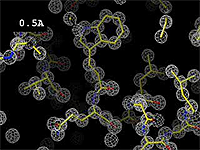
The images at right show how the electron density map[1] becomes more accurate and detailed as the uncertainty (resolution value) decreases from 5.0 Å to 0.5 Å.
See Also
Websites
- Resolution at ProteinExplorer.Org's Glossary.
Notes
- ↑ These are "perfect" electron density maps calculated from the atomic model (R factor = 0.0%, perfect phases and amplitudes, contoured at 1 sigma). Electron density maps based on experimental data would fit the true conformation less well. Because these electron density maps were calculated from an atomic model that lacked hydrogen atoms, the electron densities for hydrogen atoms that would appear with experimental data at a resolution of 0.5 Å do not appear.
Proteopedia Page Contributors and Editors (what is this?)
Eric Martz, Joel L. Sussman, Wayne Decatur, Eran Hodis, YongLiang Jiang, Jaime Prilusky