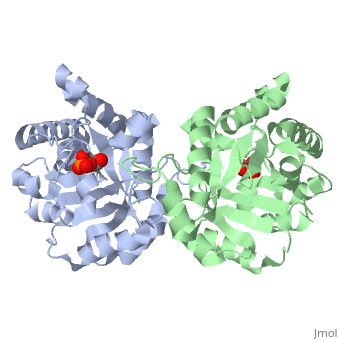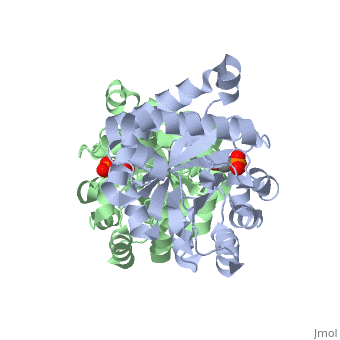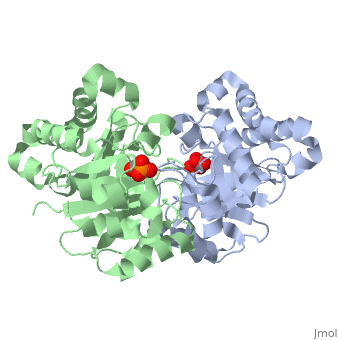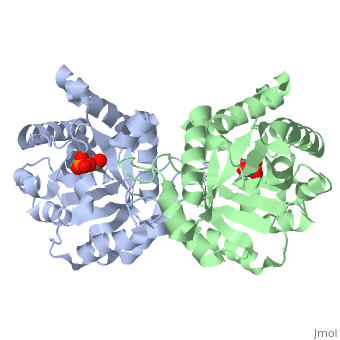This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 806
From Proteopedia
| This Sandbox is Reserved from Oct 10, 2013, through May 20, 2014 for use in the course "CHEM 410 Biochemistry 1 and 2" taught by Hanna Tims at the Messiah College. This reservation includes Sandbox Reserved 780 through Sandbox Reserved 807. |
To get started:
More help: Help:Editing |
|
Contents |
Introduction
This is Triosephosphate isomerase, an enzyme in the glycolytic pathway. It is a dimer, but can function as a . It catalyzes the reaction of D-glyceraldehyde 3-phosphate to glycerone phosphate.
3D Structure
The contains alpha helices (gold) and beta sheets (purple). Overall, it forms a beta barrel motif.
was not working.
Residues
Pictured in yellow are the . The are pictured in green.
Solvent Accessibility
are pictured in dark blue. They are mostly located between the beta sheets and alpha helices, in a diagonal line, closer to where the monomers connect to form a dimer.
Ligand Interaction Site
The ligand with the molecule at the location pictured.