User:Clayton Moore/Sandbox 1
From Proteopedia
Contents |
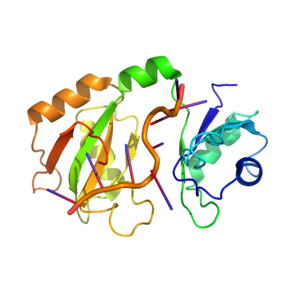
Hrp1-Saccharomyces cervisiae
|
Hrp1 is a heterogeneous ribonuclear protein of Saccharomyces cerevisiae, baker’s yeast. Hrp1 is an essential component of 3’ pre-mRNA processing and contributes to the preparatory cleavage required for polyadenylation. The gene expressed as Hrp1, HRP1, was first isolated by Henry, et al.[1] and was later attributed to the Hrp1 protein by Kessler, et al.[2] Hrp1 also participates in the regulation of the 3’ end.
Function
Polyadenylation Complex
Hrp1 is a member of yeast mRNA cleavage factor 1 (CF1), which, along with the cleavage stimulatory factor (CstF), processes the 3’ end, most notably through polyadenylation. Though crucial in eukaryotic pre-mRNA processing, polyadenylation is especially important in yeast, where intron splicing is far less frequent than in higher eukaryotes.[3] CF1 binds mRNA upstream of the cleavage site[4] and is divided into two components, CF1a and b, the former of which contains Rna14, Rna15, Clp1 and Pfc11.[5] Hrp1 is the sole polypeptide of cleavage factor 1b (CF1b).[6] Hrp1 interacts with the CF1a proteins Rna14 and Rna15 in an inverted U-like structure in the presence of mRNA.[7] Rna15 interacts with both RNA recognition motifs (RRMs) of Hrp15[8], while Rna14 contacts Hrp1 in such a way that maximizes the distance between the negative domains of each protein.[9]
Specificity and Location
Yeast hnRNPs, including Hrp1, are specific to certain mRNA strands. Hrp1 is specific to an mRNA efficiency element consisting of alternating UA sequencing[10] approximately seven nucleotides in length.[11] This sequence specificity, among other observational data, has given credence to the notion, first proposed by Minvielle-Sebastia, et al., that Hrp1 is not totally essential to the 3’ processing of every mRNA.[12] This notion is further supported by the ability of yeast cells to survive hyperosmotic stress-induced extranuclear export of Hrp1.[13]
Hrp1 participates in mRNA processing within the nucleus, but it may be found endo- or exonuclearly.[14],[15] A nuclear localization signal (NLS) at the C-terminal end of Hrp1 is essential to its recognition by the nuclear transportin receptor Kap104. The receptor and NLS are orthologous to human karyopherin B2 and hnRNP A1.[16]
Regulatory Function
Hrp1 participates in multiple regulatory pathways involving 3’ processing.[17] It has been implicated in the inhibition of transcription by the Sen-1 helicase as mediated by Nrd-1.[18] Hrp1 also activates the nonsense-mediated mRNA decay pathway, which monitors translation and degrades incorrect mRNA.[19] Hrp1, along with Rna14, competes with Npl13, which inhibits cleavage at the polyadenylation site.[20]
Disease
Though Hrp1 is not analogous to any mammalian hnRNP[21], the protein and its corresponding gene are occasionally studied as orthologues to human hnRNPs. HNRPDL is one such family of human hnRNPs. Mutations to several members of this class of hnRNPs result in many facets of muscular dystrophy. A study by Vieira, et al.[22] found that elimination of Hrp1 had profound effects on protein localization and activation, and these results were used as a model for the genotypic causation of muscular dystrophy.
References
- ↑ Henry, Michael, et al. “Potential RNA Binding Proteins in Saccharomyces Cerevisiae Identified as Suppressors of Temperature-Sensitive Mutations inNPL3.” Genetics, vol. 142, Jan. 1996, pp. 103–115.
- ↑ Kessler, Marco M, et al. “Purification of the Saccharomyces Cerevisiae Cleavage/Polyadenylation Factor I.” Journal of Biological Chemistry, vol. 271, no. 43, 25 Oct. 1996, pp. 27167–27175.
- ↑ Guisbert, K. Kim. “Functional Specificity of Shuttling HnRNPs Revealed by Genome-Wide Analysis of Their RNA Binding Profiles.” RNA, vol. 11, no. 4, Jan. 2005, pp. 383–393., doi:10.1261/rna.7234205.
- ↑ Kessler, Marco M, et al. “Purification of the Saccharomyces Cerevisiae Cleavage/Polyadenylation Factor I.” Journal of Biological Chemistry, vol. 271, no. 43, 25 Oct. 1996, pp. 27167–27175.
- ↑ Minvielle-Sebastia, L. “Control of Cleavage Site Selection during MRNA 3 End Formation by a Yeast HnRNP.” The EMBO Journal, vol. 17, no. 24, 1998, pp. 7454–7468., doi:10.1093/emboj/17.24.7454.
- ↑ Kessler, M. M., et al. “Hrp1, a Sequence-Specific RNA-Binding Protein That Shuttles between the Nucleus and the Cytoplasm, Is Required for MRNA 3-End Formation in Yeast.” Genes & Development, vol. 11, no. 19, Jan. 1997, pp. 2545–2556., doi:10.1101/gad.11.19.2545.
- ↑ Barnwal, R. P., et al. “Structural and Biochemical Analysis of the Assembly and Function of the Yeast Pre-MRNA 3 End Processing Complex CF I.” Proceedings of the National Academy of Sciences, vol. 109, no. 52, Oct. 2012, pp. 21342–21347., doi:10.1073/pnas.1214102110.
- ↑ Barnwal, R. P., et al. “Structural and Biochemical Analysis of the Assembly and Function of the Yeast Pre-MRNA 3 End Processing Complex CF I.” Proceedings of the National Academy of Sciences, vol. 109, no. 52, Oct. 2012, pp. 21342–21347., doi:10.1073/pnas.1214102110.
- ↑ Leeper, Thomas C., et al. “Novel Protein–Protein Contacts Facilitate MRNA 3′-Processing Signal Recognition by Rna15 and Hrp1.” Journal of Molecular Biology, vol. 401, no. 3, 2010, pp. 334–349., doi:10.1016/j.jmb.2010.06.032.
- ↑ Guisbert, K. Kim. “Functional Specificity of Shuttling HnRNPs Revealed by Genome-Wide Analysis of Their RNA Binding Profiles.” RNA, vol. 11, no. 4, Jan. 2005, pp. 383–393., doi:10.1261/rna.7234205.
- ↑ Chen, S. “A Specific RNA-Protein Interaction at Yeast Polyadenylation Efficiency Elements.” Nucleic Acids Research, vol. 26, no. 21, Jan. 1998, pp. 4965–4974., doi:10.1093/nar/26.21.4965.
- ↑ Minvielle-Sebastia, L. “Control of Cleavage Site Selection during MRNA 3 End Formation by a Yeast HnRNP.” The EMBO Journal, vol. 17, no. 24, 1998, pp. 7454–7468., doi:10.1093/emboj/17.24.7454.
- ↑ Henry, M. F. “The Yeast HnRNP-like Protein Hrp1/Nab4 Accumulates in the Cytoplasm after Hyperosmotic Stress: A Novel Fps1-Dependent Response.” Molecular Biology of the Cell, vol. 14, no. 9, Nov. 2003, pp. 3929–3941., doi:10.1091/mbc.e03-01-0854.
- ↑ Henry, M. F. “The Yeast HnRNP-like Protein Hrp1/Nab4 Accumulates in the Cytoplasm after Hyperosmotic Stress: A Novel Fps1-Dependent Response.” Molecular Biology of the Cell, vol. 14, no. 9, Nov. 2003, pp. 3929–3941., doi:10.1091/mbc.e03-01-0854.
- ↑ Kessler, M. M., et al. “Hrp1, a Sequence-Specific RNA-Binding Protein That Shuttles between the Nucleus and the Cytoplasm, Is Required for MRNA 3-End Formation in Yeast.” Genes & Development, vol. 11, no. 19, Jan. 1997, pp. 2545–2556., doi:10.1101/gad.11.19.2545.
- ↑ Lange, Allison, et al. “A PY-NLS Nuclear Targeting Signal Is Required for Nuclear Localization and Function of TheSaccharomyces CerevisiaemRNA-Binding Protein Hrp1.” Journal of Biological Chemistry, vol. 283, no. 19, 2008, pp. 12926–12934., doi:10.1074/jbc.m800898200.
- ↑ Tuck, Alex C, and David Tollervey. “A Transcriptome-Wide Atlas of RNP Composition Reveals Diverse Classes of MRNAs and LncRNAs.” Cell, vol. 154, 29 Aug. 2013, pp. 996–1009.
- ↑ Kuehner, Jason N., and David A. Brow. “Regulation of a Eukaryotic Gene by GTP-Dependent Start Site Selection and Transcription Attenuation.” Molecular Cell, vol. 31, no. 2, 2008, pp. 201–211., doi:10.1016/j.molcel.2008.05.018.
- ↑ González, Carlos I., et al. “The Yeast HnRNP-like Protein Hrp1/Nab4 Marks a Transcript for Nonsense-Mediated MRNA Decay.” Molecular Cell, vol. 5, no. 3, 2000, pp. 489–499., doi:10.1016/s1097-2765(00)80443-8.
- ↑ Bucheli, M. E., et al. “Polyadenylation Site Choice in Yeast Is Affected by Competition between Npl3 and Polyadenylation Factor CFI.” RNA, vol. 13, no. 10, 2007, pp. 1756–1764., doi:10.1261/rna.607207.
- ↑ Gross, S., and C. Moore. “Five Subunits Are Required for Reconstitution of the Cleavage and Polyadenylation Activities of Saccharomyces Cerevisiae Cleavage Factor I.” Proceedings of the National Academy of Sciences, vol. 98, no. 11, Aug. 2001, pp. 6080–6085., doi:10.1073/pnas.101046598.
- ↑ Vieira, Natássia M., et al. “A Defect in the RNA-Processing Protein HNRPDL Causes Limb-Girdle Muscular Dystrophy 1G (LGMD1G).” Human Molecular Genetics, vol. 23, no. 15, 2014, pp. 4103–4110., doi:10.1093/hmg/ddu127.