This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Molecular Playground/1,3-1,4-beta-D-glucanase
From Proteopedia
1,3-1,4-beta-D-glucanase
Endo-1,4-beta-D-glucanases (EGases) form a large family of hydrolytic enzymes in prokaryotes and eukaryotes. In higher plants, potential substrates in vivo are xyloglucan and non-crystalline cellulose in the cell wall. Gene expression patterns suggest a role for EGases in various developmental processes such as leaf abscission, fruit ripening and cell expansion.
| |||||||
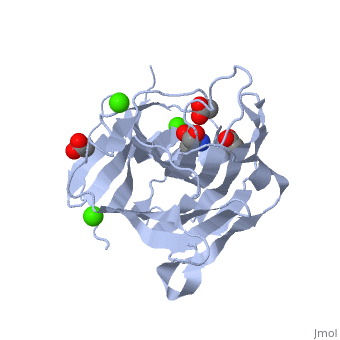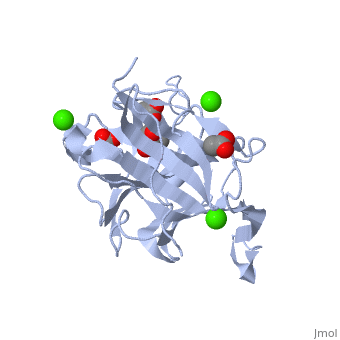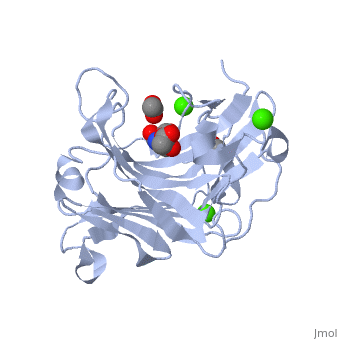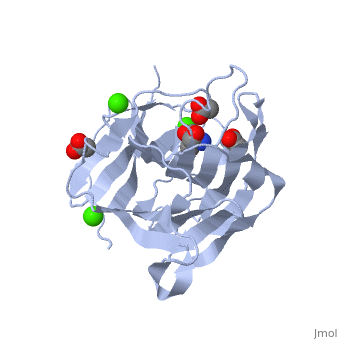
| β-glucanase complex with Tris, acetate and Ca+2 ion, 3h0o | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | , , | ||||||
| Activity: | Licheninase, with EC number 3.2.1.73 | ||||||
| Related: | 1mve, 1zm1, 3hr9 | ||||||
| |||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||