This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Glucanase
From Proteopedia
(Difference between revisions)
| Line 7: | Line 7: | ||
Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
{{#tree:id=OrganizedByTopic|openlevels=0| | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
| + | |||
| + | *Endo-1,2–β-glucanase | ||
| + | |||
| + | **[[5gzh]] - Endo-Gase – ''Chitinophaga linensis''<br /> | ||
*Endo-1,3-β-glucanase | *Endo-1,3-β-glucanase | ||
| Line 24: | Line 28: | ||
**[[5fui]] - ZgEndo-Gase C terminal <br /> | **[[5fui]] - ZgEndo-Gase C terminal <br /> | ||
**[[4pq9]] - Endo-Gase – ''Mycobacterium marinum'' <br /> | **[[4pq9]] - Endo-Gase – ''Mycobacterium marinum'' <br /> | ||
| + | **[[5h9x]] - PbEndo-Gase – ''Paenibacillus barengoltzii'' <br /> | ||
| + | **[[4wzf]] - MtEndo-Gase – ''Mycobacterium tuberculosis''<br /> | ||
*''Endo-1,3– β-glucanase binary complex'' | *''Endo-1,3– β-glucanase binary complex'' | ||
**[[4gzi]], [[4gzj]] - pEndo-Gase (mutant) + glucose<br /> | **[[4gzi]], [[4gzj]] - pEndo-Gase (mutant) + glucose<br /> | ||
| - | **[[4bpz]] - ZgEndo-Gase catalytic domain (mutant) + trisasccharide<br /> | + | **[[4bpz]], [[4cte]] - ZgEndo-Gase catalytic domain (mutant) + trisasccharide<br /> |
| + | **[[5h9y]] - PbEndo-Gase + laminarihexaose <br /> | ||
*Endo-1,4-β-glucanase | *Endo-1,4-β-glucanase | ||
| Line 37: | Line 44: | ||
**[[1j83]] - CcEndo-Gase ENGF <br /> | **[[1j83]] - CcEndo-Gase ENGF <br /> | ||
**[[3ik2]] - Endo-Gase A residues 34-541 – ''Clostridium acetobulylicum''<br /> | **[[3ik2]] - Endo-Gase A residues 34-541 – ''Clostridium acetobulylicum''<br /> | ||
| - | **[[2xfg]] - CtEndo-Gase 1 catalytic and cellulose-binding domains – ''Clostridium thermocellum''<br /> | + | **[[2xfg]], [[5byw]] - CtEndo-Gase 1 catalytic and cellulose-binding domains – ''Clostridium thermocellum''<br /> |
**[[1cem]], [[2e0p]], [[1is9]] - CtEndo-Gase A catalytic domain<br /> | **[[1cem]], [[2e0p]], [[1is9]] - CtEndo-Gase A catalytic domain<br /> | ||
**[[2wab]] - CtEndo-Gase E C terminal domain (mutant) <br /> | **[[2wab]] - CtEndo-Gase E C terminal domain (mutant) <br /> | ||
| Line 88: | Line 95: | ||
**[[4v2x]] - BhEndo-Gase – ''Bacillus halodurans'' <br /> | **[[4v2x]] - BhEndo-Gase – ''Bacillus halodurans'' <br /> | ||
**[[4uz8]], [[4uzn]] - BhEndo-Gase carbohydrate-binding module <br /> | **[[4uz8]], [[4uzn]] - BhEndo-Gase carbohydrate-binding module <br /> | ||
| + | **[[5i77]] - AnEndo-Gase – ''Aspergillus niger''<br /> | ||
| + | **[[5i78]] - AnEndo-Gase (mutant)<br /> | ||
| + | **[[5h4u]] - Endo-Gase – ''Cryptopygus antarcticus'' <br /> | ||
| + | **[[5czl]] - Endo-Gase – ''Raoultella ornithinolytica'' <br /> | ||
| + | **[[5cd2]] - Endo-Gase – ''Vibrio fischeri'' <br /> | ||
*''Endo-1,4–β-glucanase binary complex'' | *''Endo-1,4–β-glucanase binary complex'' | ||
| Line 120: | Line 132: | ||
**[[3wdu]], [[3wdv]] - PtEndo-Gase + polysaccharide<br /> | **[[3wdu]], [[3wdv]] - PtEndo-Gase + polysaccharide<br /> | ||
**[[3wdx]], [[3wdy]] - PtEndo-Gase (mutant) + polysaccharide<br /> | **[[3wdx]], [[3wdy]] - PtEndo-Gase (mutant) + polysaccharide<br /> | ||
| + | **[[5i79]] - AnEndo-Gase (mutant) + polysaccharide<br /> | ||
*Endo-1,3-1,4-β-glucanase | *Endo-1,3-1,4-β-glucanase | ||
| Line 132: | Line 145: | ||
**[[1glh]] - Endo-Gase – synthetic<br /> | **[[1glh]] - Endo-Gase – synthetic<br /> | ||
**[[1ghr]], [[1ghs]] - Endo-Gase – ''Hordeum vulgare''<br /> | **[[1ghr]], [[1ghs]] - Endo-Gase – ''Hordeum vulgare''<br /> | ||
| - | **[[1mac]] – Gase – ''Paenibacillus macerans''<br /> | + | **[[1mac]], [[5xd0]] – Gase – ''Paenibacillus macerans''<br /> |
**[[3wdt]] – PtEndo-Gase – ''Paecilomyces thermophila''<br /> | **[[3wdt]] – PtEndo-Gase – ''Paecilomyces thermophila''<br /> | ||
**[[3wdw]] – PtEndo-Gase (mutant)<br /> | **[[3wdw]] – PtEndo-Gase (mutant)<br /> | ||
| + | **[[4x0v]] - CbEndo-Gase GH5 module <br /> | ||
*''Endo-1,3-1,4–β-glucanase binary complex'' | *''Endo-1,3-1,4–β-glucanase binary complex'' | ||
| Line 160: | Line 174: | ||
**[[1tvn]] - PhEndo-Gase G catalytic domain – ''Pseudoalteromonas haloplanktis''<br /> | **[[1tvn]] - PhEndo-Gase G catalytic domain – ''Pseudoalteromonas haloplanktis''<br /> | ||
**[[1l8f]] - MaEndo-Gase – ''Melanocarpus albomyces''<br /> | **[[1l8f]] - MaEndo-Gase – ''Melanocarpus albomyces''<br /> | ||
| - | **[[1ks4]], [[1ks5]] - | + | **[[1ks4]], [[1ks5]] - AnEndo-Gase A <br /> |
**[[1gzj]] - Endo-Gase – ''Thermoascus aurantiacus''<br /> | **[[1gzj]] - Endo-Gase – ''Thermoascus aurantiacus''<br /> | ||
**[[1e8p]], [[1e8q]], [[2j4m]], [[2j4n]] - PeEndo-Gase cellulose docking domain – ''Piromyces'' – NMR<BR /> | **[[1e8p]], [[1e8q]], [[2j4m]], [[2j4n]] - PeEndo-Gase cellulose docking domain – ''Piromyces'' – NMR<BR /> | ||
| Line 178: | Line 192: | ||
**[[4ba6]] - EucEndo-Gase Cel5a C terminal<br /> | **[[4ba6]] - EucEndo-Gase Cel5a C terminal<br /> | ||
**[[4aek]], [[4aem]] - EucEndo-Gase carbohydrate-binding domain<br /> | **[[4aek]], [[4aem]] - EucEndo-Gase carbohydrate-binding domain<br /> | ||
| + | **[[5gy3]] - Endo-Gase – ''Klebsiella pneumoniae''<br /> | ||
| + | **[[4xeb]] - Endo-Gase – ''Penicillium funiculosum'' <br /> | ||
| + | **[[4tuf]] - Endo-Gase – ''Xanthomonas campestris'' <br /> | ||
*''Non-specific endo-glucanase binary complex'' | *''Non-specific endo-glucanase binary complex'' | ||
| Line 191: | Line 208: | ||
**[[2ovw]], [[4ovw]], [[1ovw]] - FoEndo-Gase I + polysaccharide<br /> | **[[2ovw]], [[4ovw]], [[1ovw]] - FoEndo-Gase I + polysaccharide<br /> | ||
**[[3eng]], [[4eng]] - HiEndo-Gase V catalytic domain + polysaccharide<br /> | **[[3eng]], [[4eng]] - HiEndo-Gase V catalytic domain + polysaccharide<br /> | ||
| - | **[[1uoz]], [[1up0]], [[1up2]], [[1up3]] - | + | **[[1uoz]], [[1up0]], [[1up2]], [[1up3]] - MtEndo-Gase + polysaccharide <br /> |
**[[2jeq]] - PpEndo-Gase + polysaccharide<br /> | **[[2jeq]] - PpEndo-Gase + polysaccharide<br /> | ||
**[[2zey]] - TpEndo-Gase + polysaccharide<br /> | **[[2zey]] - TpEndo-Gase + polysaccharide<br /> | ||
| Line 197: | Line 214: | ||
**[[3oea]], [[3oeb]] - Endo-Gase (mutant) + polysaccharide – ''Caldanaerobius polysaccharolyticus''<br /> | **[[3oea]], [[3oeb]] - Endo-Gase (mutant) + polysaccharide – ''Caldanaerobius polysaccharolyticus''<br /> | ||
**[[3nco]] - FnEndo-Gase + peptide<br /> | **[[3nco]] - FnEndo-Gase + peptide<br /> | ||
| - | **[[4afd]] - Endo-Gase Cel5a N terminal + polysaccharide | + | **[[4afd]] - Endo-Gase Cel5a N terminal + polysaccharide<br /> |
| + | **[[5dzg]], [[5dze]], [[5dzf]] - Endo-Gase (mutant) + polysaccharide – grape<br /> | ||
*Exo-1,3-β-glucanase | *Exo-1,3-β-glucanase | ||
Revision as of 10:23, 24 May 2017
| |||||||||||
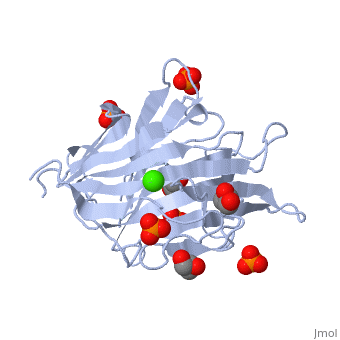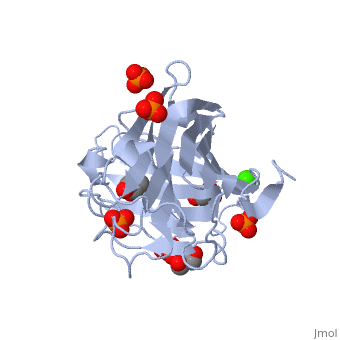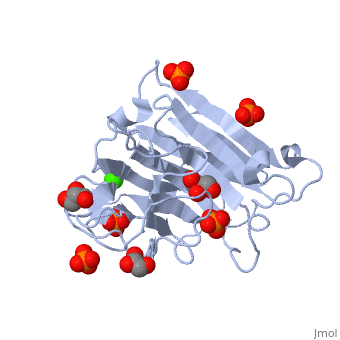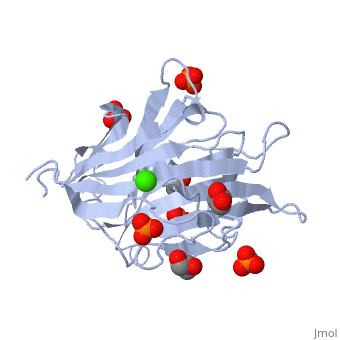
3D structures of glucanase
Updated on 24-May-2017
3fw6, 3ii1, 4ee9, 4m24 - ubEndo-Gase – uncultured bacterium
References
- ↑ Barras DR, Stone BA. Beta-1,3-glucan hydrolases from Euglena gracilis. I. The nature of the hydrolases. Biochim Biophys Acta. 1969 Nov 4;191(2):329-41. PMID:5354264
- ↑ Tsai LC, Amiraslanov I, Chen HR, Chen YW, Lee HL, Liang PH, Liaw YC. Structures of exoglucanase from Clostridium cellulovorans: cellotetraose binding and cleavage. Acta Crystallogr F Struct Biol Commun. 2015 Oct 1;71(Pt 10):1264-72. doi:, 10.1107/S2053230X15015915. Epub 2015 Sep 23. PMID:26457517 doi:http://dx.doi.org/10.1107/S2053230X15015915
- ↑ Vincken JP, Beldman G, Voragen AG. Substrate specificity of endoglucanases: what determines xyloglucanase activity? Carbohydr Res. 1997 Mar 13;298(4):299-310. PMID:9098958